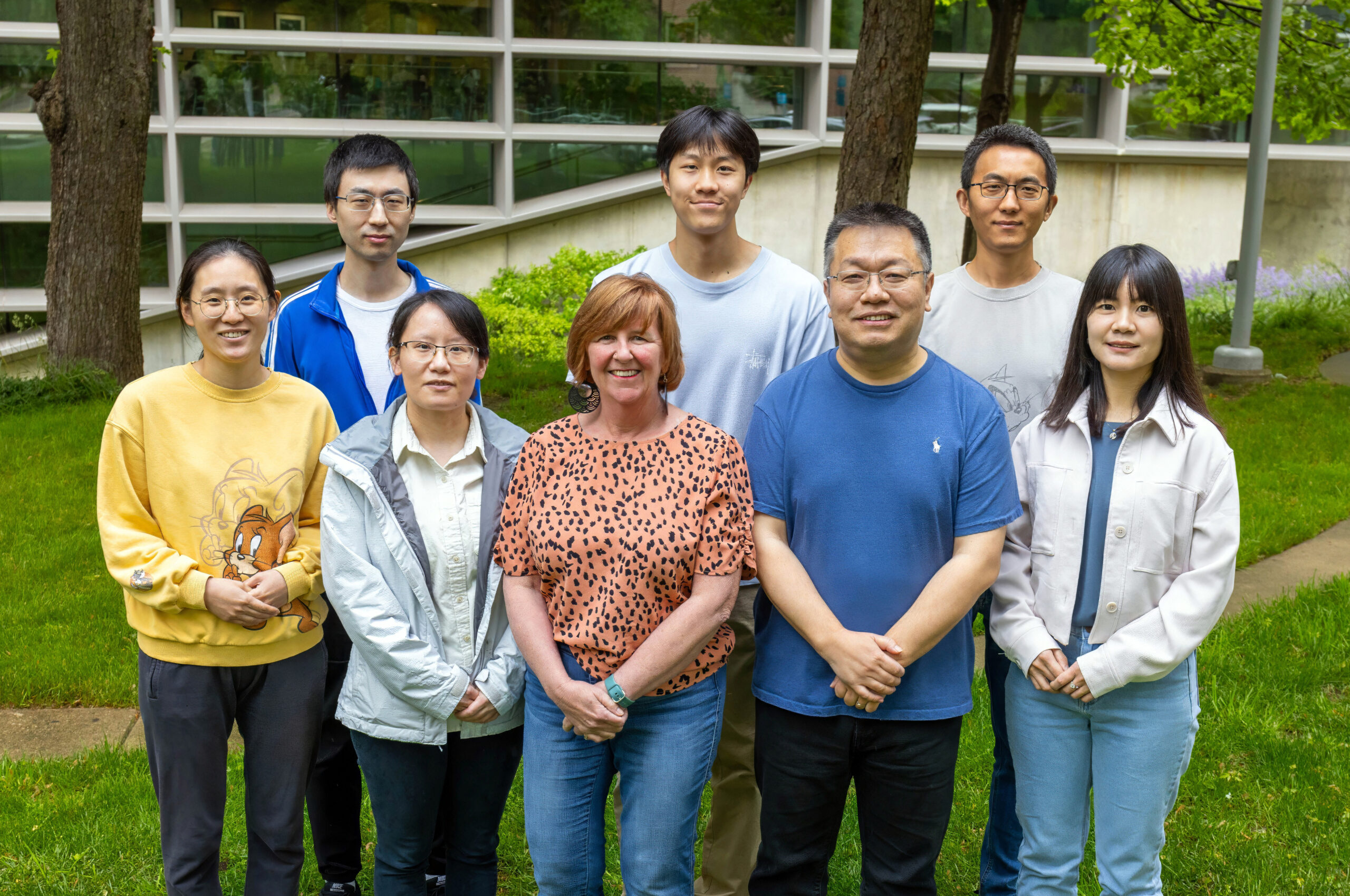
Shi Laboratory
 Histone Modifications and Chromatin Regulation
Histone Modifications and Chromatin Regulation
Each year, more than 1.6 million people in the U.S. are diagnosed with cancer. Globally, the number of people with the disease is expected to rise 50 percent in the coming years, jumping from 14 million in 2012 to an estimated 21 million by 2030.
This impending increase reinforces the importance of understanding the complex constellation of factors that contribute to cancer and leveraging this knowledge to find new, improved prevention and treatment methods. To this end, the Shi Laboratory seeks to discern how epigenetics — mechanisms that regulate how DNA is read and acted upon — impact cancer risk, development and progression.
Specifically, the Shi Laboratory investigates a number of posttranslational modifications on histones, which play an important role in modulating chromatin dynamics and the accessibility of the underlying DNA, thus regulating all chromatin-associated processes, such as transcription. Dysregulation of histone modification homeostasis leads to pathogenesis of developmental disorders and genetic diseases, including cancer.
However, despite the clear biological and clinical importance, it remains largely unknown how modifications on the histone tails dictate gene expression profiles. One of the principal functions of histone methylation and acetylation is believed to be to recruit or repel “reader” proteins that recognize the methyl or acetyl moiety on specific residues and transduce the epigenetic signals to distinct outcomes. Therefore, identification of reader proteins is critical for understanding the mechanistic and functional consequences of histone modifications.
The long-term goal of the Shi Laboratory’s research is to understand the molecular mechanisms by which histone modifications, particularly lysine methylation and acetylation, regulate chromatin and transcription and how dysregulation of the homeostasis of histone modifications leads to human cancer. Their current research focus is to identify and characterize novel epigenetic readers and to elucidate their roles in cancer.
Featured News & Recent Publications
Learn More
New anti-cancer ‘degrader’ targets protein essential to infant leukemia

Targeting uncontrolled inflammation may hold the key to treating therapy-resistant cancers

Engineering a lasting legacy: Prein&Newhof and Thomas and Garretta Newhof
Compton SE, Kitchen-Goosen SM, DeCamp LM, Lau KH, Mabvakure, Vos M, Williams KS, Wong KK, Shi X, Rothbart SB, Krawczyk CK, Jones RG. 2023. LKB1 controls inflammatory potential through CRTC2-dependent histone acetylation. Mol Cell 83(11):1872–1886.
*Featured as the cover of Molecular Cell
Wan L et al. ENL links histone acetylation to oncogenic gene expression in AML. Nature
Our Impact
We’re raising thousands to save millions.
We’re turning hope into action for the millions of people around the world affected by diseases like cancer and Parkinson’s. Find out how you can help us make a difference.
- 141 peer-reviewed papers published in 2025, 74 of which were in high-impact journals
- 15 VAI-SU2C Epigenetics Dream Team clinical trials launched to date
- 10 clinical trials and related projects supported by VAI through the International Linked Clinical Trials Program
Xiaobing Shi, Ph.D.
Professor, Department of Epigenetics
Areas of Expertise
Chromatin, post-translational modifications, epigenetics, cancer
Biography
Dr. Xiaobing Shi is an expert in cancer epigenetics and a professor in the Department of Epigenetics at Van Andel Institute. He earned a B.S. in biology from Wuhan University followed by a Ph.D. in biology from the Chinese Academy of Sciences, both with honors. He subsequently completed postdoctoral fellowships in the labs of Drs. Arthur Kornberg and Or Gozani at Stanford University before accepting a faculty position at University of Texas MD Anderson Cancer Center, where he most recently served as an Associate Professor and Co-Director of the Center for Cancer Epigenetics. Dr. Shi’s work has led to the discovery of several new readers of histone posttranslational modifications, particularly in acetylation and methylation, and elucidated their impact on cancer development. He is the recipient of several prestigious awards, including a Career Development Award from the Leukemia & Lymphoma Society, a Research Scholar Award from the American Cancer Society, a Kimmel Scholar Award from the Sidney Kimmel Foundation, and an inaugural R. Lee Clark Fellow from UT M.D. Anderson Cancer Center.
Society Memberships
American Society for Biochemistry and Molecular Biology
American Association for Cancer Research
American Association for the Advancement of Science
2025
Becht DC, Selvam K, Lachance C, Côté V, Li K, Nguyen MC, Pareek A, Shi X, Wen H, Blanco MA, Côté J, Kutateladze TG. 2025. A multivalent engagement of ENL with MOZ. Nat Struct Mol Biol 32(4):709–718.
Vann KR, Sharma R, Hsu CC, Devoucoux M, Tencer AH, Zeng L, Lin K, Zhu L, Li Q, Lachance C, Ospina RR, Tong Q, Cheung KL, Yang S, Biswas S, Xuan H, Gatchalian J, Alamillo L, Wang J, Jang SM, Klein BJ, Lu Y, Ernst P, Strahl BD, Rothbart SB, Walsh MJ, Cleary ML, Côté J, Shi X, Zhou MM, Kutateladze TG. 2025. Structure-function relationship of ASH1L and histone H3K36 and H3K4 methylation. Nat Commun 16(1):223
Xue Z, Xuan H, Lau K, Su Y, Wegener M, Li K, Turner L, Adams M, Shi X, Wen H. 2025. Expression of ENL YEATS domain tumor mutations in nephrogenic or stromal lineage impairs kidney development. Nat Commun 16:2531.
2024
Xu L, Xuan H, Shi X. 2025. Dysregulation of the p300/CBP histone acetyltransferases in human cancer. Epigenomics 17(3):193–208.
Wang H, Wen H, Shi X. 2024. The MYC-MAF-SAGA axis drives oncogenic gene expression in multiple myeloma. Genes Dev 38(15–16):693–694.
Xue Z, Qin L, Xuan H, Luo K, Huang M, Xie L, Su Y, Xu L, Harsh J, Dale B, Shi X, Chen X, Kaniskan JÜ, Jin J*, Wen H*. 2024. A potent and selective ENL degrader suppresses oncogenic gene expression and leukemia progression. Sci Adv 10(35).
Xuan H, Xu L, Li K, Xuan F, Xu T, Wen H, Shi X. 2024. Hotspot cancer mutation impairs KAT8-mediated nucleosomal histone acetylation. J Mol Biol 436(7):168413.
Xuan F, Xuan H, Huang M, He W, Xu H, Shi X, Wen H. 2024. The Tudor-knot domain of KAT5 regulates nucleosomal substrate acetylation. J Mol Biol 436(7):168414.
Kumar Sinha V, Zhang Y, Xu L, Chen YW, Picaud S, Zandian M, Biswas S, Filippakopoulos P, Wang SP, Shi X, Kutateladze TG. 2024. Histone H4K16ac binding function of the triple PHD finger cassette of MLL4. J Mol Biol 436(7):168212.
Becht DC, Kanai A, Biswas S, Halawa M, Zeng L, Cox KL, Poirier MG, Zhou MM, Shi X, Yokoyama A, Kutateladze TG. 2024. The winged helix domain of MORF binds CpG islands and the TAZ2 domain of p300. iScience 27(4):109367.
2023
Wen H, Shi X. 2023. Histone readers and their roles in cancer. Cancer Treat Res 190:245–272.
Xu L*, Xuan H*, He W, Zhang L, Huang M, Li K, Wen H, Xu H, Shi X. 2023. TAZ2 truncation confers overactivation of p300 and cellular vulnerability to HDAC inhibition. Nat Commun. 14(1):5362.
Tencer AH*, Yu Y*, Causse SZ*, Campbell GR, Klein BJ, Xuan H, Cartier J, Miles MA, Gaurav N, Zadoroznyj A, Holt TA, Wen H, Hawkins CJ, Spector SA, Dubrez L*, Shi X*, Kutateladze TG*. 2023. Molecular basis for nuclear accumulation and targeting of the inhibitor of apoptosis BIRC2. Nat Struct Mol Biol. 30(9):1265-1274.
Kumar Sinha V, Zhang Y, Xu L, Chen YW, Picaud S, Zandian M, Biswas S, Filippakopoulos P, Wang SP, Shi X, Kutateladze TG. 2023. Histone H4K16ac Binding Function of the Triple PHD Finger Cassette of MLL4. J Mol Biol. 20:168212.
Compton SE, Kitchen-Goosen SM, DeCamp LM, Lau KH, Mabvakure, Vos M, Williams KS, Wong KK, Shi X, Rothbart SB, Krawczyk CK, Jones RG. 2023. LKB1 controls inflammatory potential through CRTC2-dependent histone acetylation. Mol Cell. 83(11):1872-1886.
TRIM28 secures skeletal stem cell fate during skeletogenesis by silencing neural gene expression and repressing GREM1/AKT/mTOR signaling axis. Cell Rep. 42(1):112012.
2022
Liu X, Wang J, Boyer JA, Gong W, Zhao S, Xie L, Wu Q, Zhang C, Jain K, Guo Y, Rodriguez J, Li M, Uryu H, Liao C, Zhou, Shi X, Tsai Y-H, Yan Q, Lou W, Chen X, Strahl BD, von Kriegsheim A, Zhang Q, Wang GG, Baldwin AS, Zhang Q. 2022. Histone H3 proline 16 hydroxylation regulates mammalian gene expression. Nature Genet. 54(11):1721–1735.
Kim KB, Kabra A, Kim DW, Xue Y, Huang Y, Hou PC, Zhou Y, Miranda LJ, Park JI, Shi X, Bender TP, Bushweller JH, Park KS. 2022. KIX domain determines a selective tumor-promoting role for EP300 and its vulnerability in small cell lung cancer. Sci Adv. 8(7):eabl4618.
2021
Zhang Y, Brown K, Yu Y, Ibrahim Z, Zandian M, Xuan H, Ingersoll S, Lee T, Ebmeier CC, Liu J, Panne D, Shi X, Ren X, Kutateladze TG. 2021. Nuclear condensates of p300 formed though the structured catalytic core can act as a storage pool of p300 with reduced HAT activity. Nat Commun 12(1):4618.
Klein BJ, Deshpande A, Cox KL, Xuan F, Zandian M, Barbosa K, Khanal S, Tong Q, Zhang Y, Zhang P, Sinha A, Bohlander SK, Shi X, Wen H, Poirier MG, Deshpande AJ, Kutateladze TG. 2021. The role of the PZP domain of AF10 in acute leukemia driven by AF10 translocations. Nat Commun 12(1):4130.
Ma XR, Xu L, Xu S, Klein BJ, Wang H, Das S, Li K, Yang KS, Sohail S, Chapman A, Kutateladze TG, Shi X, Liu WR, Wen H. 2021. Discovery of Selective Small-Molecule Inhibitors for the ENL YEATS Domain. J Med Chem 64(15):10997-11013.
Yu Y, Wen H, Shi X. 2021. Histone mimics: more tales to read. Biochem J 478(14):2789-2791.
Nakamura N, Shi X, Darabi R, Li Y. 2021. Hypoxia in cell reprogramming and the epigenetic regulations. Front Cell Dev Bio 9: 609984.
Yu Y*, Tencer A*, Xuan H*, Kutateladze TG, Shi X. 2021. ZZEF1 is a histone reader and transcriptional coregulator of Krüppel-like factors. J Mol Biol 433(2):166722.
Ren X, Zhou Y, Xue Z, Hao N, Li Y, Guo X, Wang D, Shi X, Li H. 2021. Histone benzoylation serves as an epigenetic mark for DPF and YEATS family proteins. Nucleic Acids Res 49(1):114–126.
2020
Wen H, Shi X. 2020. H3.3S31 phosphorylation: linking transcription elongation to stimulation responses. Signal Transduct Target Ther 5(1):176.
Shen X, Wang R, Kim MJ, Hu Q, Hsu CC, Yao J, Klages-Mundt N, Tian Y, Lynn E, Brewer TF, Zhang Y, Arun B, Gan B, Andreeff M, Takeda S, Chen J, Park JI, Shi X, Chang CJ, Jung SY, Qin J, Li L. 2020. A surge of DNA damage links transcriptional reprogramming and hematopoietic deficit in Fanconi anemia. Mol Cell 80(6):1013–1024.
Zhang Y, Guo Y, Gough SM, Zhang J, Vann KR, Li K, Cai L, Shi X, Aplan PD, Wang GG, Kutateladze TG. 2020. Mechanistic insights into chromatin targeting by leukemic NUP98-PHF23 fusion. Nat Commun 11(1):3339.
Liu J, Xue Z, Vann KR, Shi X, Kutateladze TG. 2020. Protocol for biochemical analysis and structure determination of the ZZ domain of the E3 ubiquitin ligase HERC2. STAR Protoc 1(3):100155.
Liu J*, Xue Z*, Zhang Y, Vann KR, Shi X*, Kutateladze TG*. 2020. Structural insight into binding of the ZZ domain of HERC2 to histone H3 and SUMO1. Structure 28(11):1225–1230.
Chen Y, Fang R, Yue C, Chang G, Li P, Guo Q, Wang J, Zhou A, Zhang S, Fuller GN, Shi X, Huang S. 2020. Wnt-induced stabilization of KDM4C is required for Wnt/β-Catenin target gene expression and glioblastoma tumorigenesis. Cancer Res 80(5):1049–1063.
2019
Klein BJ, Jang SM, Lachance C, Mi W, Lyu J, Sakuraba S, Krajewski K, Wang WW, Sidoli S, Liu J, Zhang Y, Wang X, Warfield BM, Kueh AJ, Voss AK, Thomas T, Garcia BA, Liu WR, Strahl BD, Kono H, Li W, Shi X, Côté J, Kutateladze TG. 2019. Histone H3K23-specific acetylation by MORF is coupled to H3K14 acylation. Nat Commun 10(1):4724.
He W, Zhang L, Villarreal OD, Fu R, Bedford E, Dou J, Patel AY, Bedford MT, Shi X, Chen T, Bartholomew B, Xu H. 2019. De novo identification of essential protein domains from CRISPR-Cas9 tiling-sgRNA knockout screens. Nat Commun 10(1):4541.
Gao G, Zhang L, Villarreal OD, He W, Su D, Bedford E, Moh P, Shen J, Shi X, Bedford MT, Xu H. 2019. PRMT1 loss sensitizes cells to PRMT5 inhibition. Nucleic Acids Res.
Zhang Y, Jang Y, Lee JE, Ahn J, Xu L, Holden MR, Cornett EM, Krajewski K, Klein BJ, Wang SP, Dou Y, Roeder RG, Strahl BD, Rothbart SB, Shi X, Ge K, Kutateladze TG. 2019. Selective binding of the PHD6 finger of MLL4 to histone H4K16ac links MLL4 and MOF. Nat Commun 10(1):2314.
Zhang Y*, Mi W*, Xue Y*, Shi X#, Kutateladze TG#. 2019 The ZZ domain as a new epigenetic reader and a degradation signal sensor. Crit Rev Biochem Mol Biol.;54(1):1-10. Review.
*Equal contribution
2018
Mi W*, Zhang Y*, Lyu J*, Wang X, Tong Q, Peng D, Xue Y, Tencer AH, Wen H, Li W, Kutateladze TG*, Shi X*. 2018. The ZZ-type zinc finger of ZZZ3 modulates the ATAC complex-mediated histone acetylation and gene activation. Nat Commun 9(1):3759.
Zhang Y*, Xue Y*, Shi J, Ahn J, Mi W, Ali M, Wang X, Klein BJ, Wen H, Li W, Shi X*, Kutateladze TG*. 2018. The ZZ domain of p300 mediates specificity of the adjacent HAT domain for histone H3.
Nat Struct Mol Biol 25(9):841-849.
Zhu S, Zhao D, Yan L, Jiang W, Kim JS, Gu B, Liu Q, Wang R, Xia B, Zhao JC, Song G, Mi W, Wang RF, Shi X, Lam HM, Dong X, Yu J, Chen K, Cao Q. 2018. BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1. Nat Commun 9(1):500.
Shi L*, Shi J*, X S, Li W, Wen H. 2018. Histone H3.3 G34 mutations alter histone H3K36 and H3K27 methylation in cis. J Mol Biol. pii:S0022-2836(18)30248-1.
Hsu C*, Shi J*, Yuan C*, Zhao D*, Jiang S, Lyu J, Wang X, Li H, Wen H, Li W*, Shi X*. 2018. Recognition of histone acetylation by the GAS41 YEATS domain promotes H2A.Z deposition in non-small cell lung cancer. Genes Dev 32(1):58–69.
Zhu S, Zhao D, Yan L, Jiang W, Kim JS, Gu B, Liu Q, Wang R, Xia B, Zhao JC, Song G, Mi W, Wang RF, Shi X, Lam HM, Dong X, Yu J, Chen K, Cao Q. 2018. BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1. Nat Commun 9(1):500.
Franco HL, Nagari A, Malladi VS, Li W, Xi Y, Richardson D, Allton KL, Tanaka K, Li J, Murakami S, Keyomarsi K, Bedford MT, Shi X, Li W, Barton MC, Dent SYR, Kraus WL. 2018. Enhancer transcription reveals subtype-specific gene expression programs controlling breast cancer pathogenesis. Genome Res 28(2):159–170.
Xi Y*, Shi J*, Li W*, Tanaka K*, Allton KL*, Richardson D, Li J, Franco HL, Nagari A, Malladi VS, Coletta LD, Simper MS, Keyomarsi K, Shen J, Bedford MT, Shi X, Barton MC, Lee Kraus W, Li W, Dent SYR. 2018. Histone modification profiling in breast cancer cell lines highlights commonalities and differences among subtypes. BMC Genomics 19(1):150.
*Equal contribution
Hsu CC*, Zhao D*, Shi J*, Peng D, Guan H, Li Y, Huang Y, Wen H, Li W*, Li H*, Shi X*. 2018. Gas41 links histone acetylation to H2A.Z deposition and maintenance of embryonic stem cell identity. Cell Discov. 4:28.
Klein BJ, Vann KR, Andrews FH, Wang WW, Zhang J, Zhang Y, Beloglazkina AA, Mi W, Li Y, Li H, Shi X, Kutateladze AG, Strahl BD, Liu WR, Kutateladze TG. 2018. Structural insights into the π-π-π stacking mechanism and DNA-binding activity of the YEATS domain. Nat Commun. 9(1):4574.
Li X, Li XM, Jiang Y, Liu Z, Cui Y, Fung KY, van der Beelen SHE, Tian G, Wan L, Shi X, Allis CD, Li H, Li Y, Li XD. 2018. Structure-guided development of YEATS domain inhibitors by targeting π-π-π stacking. Nature Chem Biol. 14(12):1140-1149.
Zhang Y, Mun SR, Linares JF, Ahn J, Towers CG, Ji CH, Fitzwalter BE, Holden MR, Mi W, Shi X, Moscat J, Thorburn A, Diaz-Meco MT, Kwon YT, Kutateladze TG. 2018. ZZ-dependent regulation of p62/SQSTM1 in autophagy. Nat Commun. 9(1):4373.
2017
Shi L, Wen H, Shi X. 2017. The histone variant H3.3 in transcriptional regulation and human disease. J Mol Biol 429(13):1934–1945.
Mi W, Guan H, Lyu J, Zhao D, Xi Y, Jiang S, Andrews FH, Wang X, Gagea M, Wen H, Tora L, Dent SYR, Kutateladze TG, Li W*, Li H*, Shi X*. 2017. YEATS2 links histone acetylation to tumorigenesis of non-small cell lung cancer. Nat Commun 8(1):1088.
Klein BJ, Simithy J, Wang X, Ahn J, Andrews FH, Zhang Y, Côté J, Shi X, Garcia BA, Kutateladze TG. 2017. Recognition of histone H3K14 acylation by MORF. Structure 25(4):650–654.e2.
Wan L*, Wen H*, Li Y*, Lyu J, Xi Y, Hoshii T, Joseph J, Wang X, Loh Y, Souza AL, Bradner JE, Shen L, Li W, Li H, Allis CD*, Armstrong SA*, Shi X*. 2017. ENL links histone acetylation to oncogenic gene expression in AML. Nature 543(7644):265–269.
2016
Schibler A, Koutelou E, Tomida J, Wilson-Pham M, Wang L, Lu Y, Cabrera AP, Chosed RJ, Li W, Li B, Shi X, Wood RD, Dent SY. 2016. Histone H3K4 methylation regulates deactivation of the spindle assembly checkpoint through direct binding of Mad2. Genes Dev 30(10):1187–1197.
Zhao D, Guan H, Zhao S, Mi W, Wen H, Li Y, Zhao Y, Allis CD, Shi X*, Li H*. (2016) YEATS2 is a selective histone crotonylation reader. Cell Res 26(5):629–632.
Andrews FH*, Shinsky SA*, Shanle EK, Bridgers JB, Gest A, Tsun IK, Krajewski K, Shi X, Strahl BD*, Kutateladze TG*. 2016. The Taf14 YEATS domain is a reader of histone crotonylation. Nature Chem Biol 12(6):396–398.
Li Y*, Sabari BR*, Panchenko T, Wen H, Zhao D, Guan H, Wan L, Huang H, Tang Z, Zhao Y, Roeder RG, Shi X, Allis CD*, Li H*. 2016. Molecular coupling of histone crotonylation and active transcription by AF9 YEATS domain. Mol Cell 62(2):181–193.
Zhang X, Peng D, Xi Y, Yuan C, Sagum CA, Klein BJ, Tanaka K, Wen H, Kutateladze TG, Li W, Bedford MT, Shi X. 2016. G9a-mediated methylation of ERα links the PHF20/MOF histone acetyltransferase complex to hormonal gene expression. Nature Commun 7:10810.
Klein BJ*, Wang X*, Cui G, Yuan C, Botuyan MV, Lin K, Lu Y, Wang X, Zhao Y, Bruns CJ, Mer G, Shi X*, Kutateladze TG*. 2016. PHF20 readers link methylation of histone H3K4 and p53 with H4K16 acetylation. Cell Rep 17(4):1158–1170.
Shi L, Wen H, Shi X. 2016. The histone variant H3.3 in transcriptional regulation and human disease. J Mol Biol 429(13):1934–1945.
Li N, Li Y, Lv J, Zheng X, Wen H, Shen H, Zhu G, Chen TY, Dhar SS, Kan PY, Wang Z, Shiekhattar R, Shi X, Lan F, Chen K, Li W, Li H, Lee MG. 2016. ZMYND8 reads the dual histone mark H3K4me1-H3K14ac to antagonize the expression of metastasis-linked genes. Mol Cell 63(3):470–484.
2015
Chen K, Chen Z, Wu D, Zhang L, Lin X, Su J, Rodriguez B, Xi Y, Xia Z, Chen X, Shi X, Wang Q, Li W. 2015. Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor-suppressor genes. Nature Genet 47(10):1149–1157.
Zhang X, Huang Y, Shi X. 2015. Emerging roles of lysine methylation on non-histone proteins. Cell Mol Life Sci 72(22):4257–4272.
2014
Li Y*, Wen H*, Xi Y, Tanaka K, Wang H, Peng D, Ren Y, Jin Q, Dent SR, Li W*, Li H*, Shi X*. 2014. AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 159(3):558–571.
Guo R*, Zheng L*, Park JW, Lv R, Chen H, Jiao F, Xu W, Mu S, Wen H, Qiu J, Wang Z, Yang P, Wu F, Hui J, Fu X, Shi X*, Shi YG*, Xing Y*, Lan F*, Shi Y*. 2014. BS69/ZMYND11 reads and connects histone H3.3 lysine 36 trimethylation-decorated chromatin to regulated pre-mRNA processing. Mol Cell 56(2):298–310.
Wen H, Li Y, Li H, Shi X. 2014. ZMYND11: An H3.3-specific reader of H3K36me3. Cell Cycle 13(14):2153–2154.
Wen H*, Li Y*, Xi Y*, Jiang S, Stratton S, Peng D, Tanaka K, Ren Y, Xia Z, Wu J, Li B, Barton MC, Li W*, Li H*, Shi X*. 2014. ZMYND11 links histone H3.3 K36 trimethylation to transcription elongation and tumor suppression. Nature 508(7495):263–268.
Jiang Y, Trescott L, Holcomb J, Zhang X, Brunzelle J, Sirinupong N, Shi X, Yang Z. 2014. Structural insights into estrogen receptor a methylation by histone methyltransferase SMYD2, a cellular event implicated in estrogen signaling regulation. J Mol Biol 426(20):3413–3425.
Klein BJ*, Piao L*, Xi Y, Rincon-Arano H, Rothbart SB, Peng D, Wen H, Larson C, Zheng X, Cortazar M, Peña PV, Mangan A, Bentley DL, Strahl BD, Groudine M, Li W, Shi X*, Kutateladze T*. 2014. KDM5B demethylase binds to its histone target and the product through distinct modules. Cell Reports 6(2):325–335.
2013
Wang J, Leung JW, Gong Z, Feng L, Shi X, Chen J. 2013. PHF6 regulates cell cycle progression by suppressing ribosomal RNA synthesis. J Biol Chem 288(5):3174–3183.
Zhang H, Ma ZY, Zeng L, Tanaka K, Zhang CJ, Ma J, Bai G, Wang P, Zhang SW, Liu ZW, Cai T, Tang K, Liu R, Shi X, He XJ, Zhu JK. 2013. DTF1 is a core component of RNA-directed DNA methylation and may assist in the recruitment of Pol IV. Proc Natl Acad Sci U S A 110(20):8290–8295.
Zhang X, Tanaka K, Yan J, Li J, Peng D, Jiang Y, Yang Z, Barton MC, Wen H, Shi X. 2013. Regulation of estrogen receptor α by histone methyltransferase SMYD2-mediated protein methylation. Proc Natl Acad Sci U S A 110(43):17284–17289.
2012
Qian W, Miki D, Zhang H, Liu Y, Zhang X, Tang K, Kan Y, La H, Li X, Li S, Zhu X, Shi X, Zhang K, Pontes O, Chen X, Liu R, Gong Z, Zhu JK. 2012. A histone acetyltransferase regulates active DNA demethylation in Arabidopsis. Science 336(6087):1445–1448.
Brien GL, Gambero G, O’Connell DJ, Jerman E, Turner SA, Egan CM, Dunne EJ, Jurgens MC, Wynne K, Piao L, Lohan AJ, Ferguson N, Shi X, Sinha KM, Loftus BJ, Cagney G, Bracken AP. 2012. Polycomb PHF19 binds H3K36me3 and recruits PRC2 and demethylase NO66 to embryonic stem cell genes during differentiation. Nat Struct Mol Biol 19(12):1273–1281.
Dhar SS, Lee SH, Kan PY, Voigt P, Ma L, Shi X, Reinberg D, Lee MG. 2012. Trans-tail regulation of MLL4-catalyzed H3K4 methylation by H4R3 symmetric dimethylation is mediated by a tandem PHD of MLL4. Genes Dev 26(24):2749–2762.
2011
Levy D, Kuo AJ, Chang Y, Schaefer U, Kitson C, Cheung P, Espejo A, Zee BM, Liu CL, Tangsombatvisit S, Tennen RI, Kuo AY, Tanjing S, Cheung R, Chua KF, Utz PJ, Shi X, Prinjha RK, Lee K, Garcia BA, Bedford MT, Tarakhovsky A, Cheng X, Gozani O. 2011. Lysine methylation of the NF-kB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-kB signaling. Nat Immunol 12(1):29–36.
Kuo AJ, Cheung P, Chen K, Zee BM, Kioi M, Lauring J, Xi Y, Park BH, Shi X, Garcia BA, Li W, Gozani O. 2011. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell 44(4):609–620.
2010
Wen H, Li J, Song T, Lu M, Kan PY, Lee MG, Sha B, Shi X. 2010. Recognition of histone H3K4 trimethylation by the PHD finger of PHF2 modulates histone demethylation. J Biol Chem 285(13):9322.
Kleine-Kohlbrecher D, Christensen J, Vandamme J, Abarrategui I, Bak M, Tommerup N, Shi X, Gozani O, Rappsilber J, Salcini AE, Helin K. 2010. A functional link between the histone demethylase PHF8 and the transcription factor ZNF711 in X-linked mental retardation. Mol Cell 38(2):1–14.
West LE, Roy S, Lachmi-Weiner K, Hayashi R, Shi X, Appella E, Kutateladze TG, Gozani O. 2010. The MBT repeats of L3MBTL1 link SET8-mediated p53 methylation at lysine 382 to target gene repression. J Biol Chem 285(48):37725–37732.
Tsai WW, Wang Z, Yiu TT, Akdemir KC, Xia W, Winter S, Tsai CY, Shi X, Schwarzer D, Plunkett W, Aronow B, Gozani O, Fischle W, Hung MC, Patel DJ, Barton MC. 2010. TRIM24 links a non-canonical histone signature to breast cancer. Nature 468(7326):927–932.
Selected publications before 2009
Shi X, Kachirskaia I, Yamaguchi H, West LE, Wen H, Wang EW, Dutta S, Appella E, Gozani O. 2007. Modulation of p53 function by SET8-mediated methylation at lysine 382. Mol Cell 27(4):636–46.
Shi X*, Kachirskaia I*, Walter KL, Kuo JH, Lake A, Davrazou F, Chan SM, Martin DG, Fingerman IM, Briggs SD, Howe L, Utz PJ, Kutateladze TG, Lugovskoy AA, Bedford MT, Gozani O. 2007. Proteome-wide analysis in Saccharomyces cerevisiae identifies several PHD fingers as novel direct and selective binding modules of histone H3 methylated at either lysine 4 or lysine 36. J Biol Chem 282(4):2450–2455.
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T, Carney D, Peña P, Lan F, Kaadige MR, Lacoste N, Cayrou C, Davrazou F, Saha A, Cairns BR, Ayer DE, Kutateladze TG, Shi Y, Côté J, Chua KF, Gozani O. 2006. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442(7098):96–99.
Peña PV, Davrazou F, Shi X, Walter KL, Verkhusha VV, Gozani O, Zhao R, Kutateladze TG. 2006. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442(7098):100–103.
Shi X, Rao NN, Kornberg A. 2004. Inorganic polyphosphate in Bacillus cereus: motility, biofilm formation, and sporulation. Proc Natl Acad Sci U S A 101(49):17061–17065.


Jennifer Brooks
Senior Administrative Assistant I, Department of Epigenetics

Mengying Huang, Ph.D.
Research Scientist, Department of Epigenetics

Hongkuan Wang, Ph.D.
Research Scientist, Department of Epigenetics
Identifying novel epigenetic readers and their inhibitors

Longxia Xu, Ph.D.
Research Scientist, Department of Epigenetics


